New Paper: Modeling with Alternate Locations in X-ray Protein Structure
11. April 2023
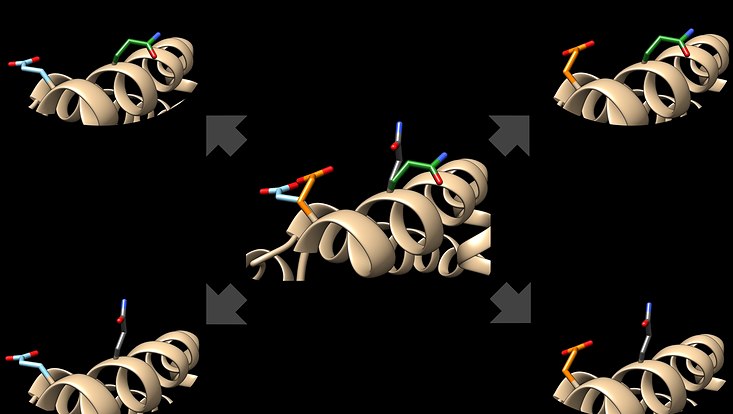
Foto: Toben Gutermuth
Crystallographic protein structures sometimes contain multiple conformations of side chains, ligands or even larger protein parts like flexible loops. The PDB format allows the annotation of alternative conformations with so-called AltLoc-Indicators, however most modeling tools do not make use of this valuable information. Our recent publication introduces a software tool to create clash-free conformations from AltLoc combinations. More about the AltLoc-Enumerator tool, its algorithm, AltLoc statistics in the PDB and our first experiences with it can be found here.